  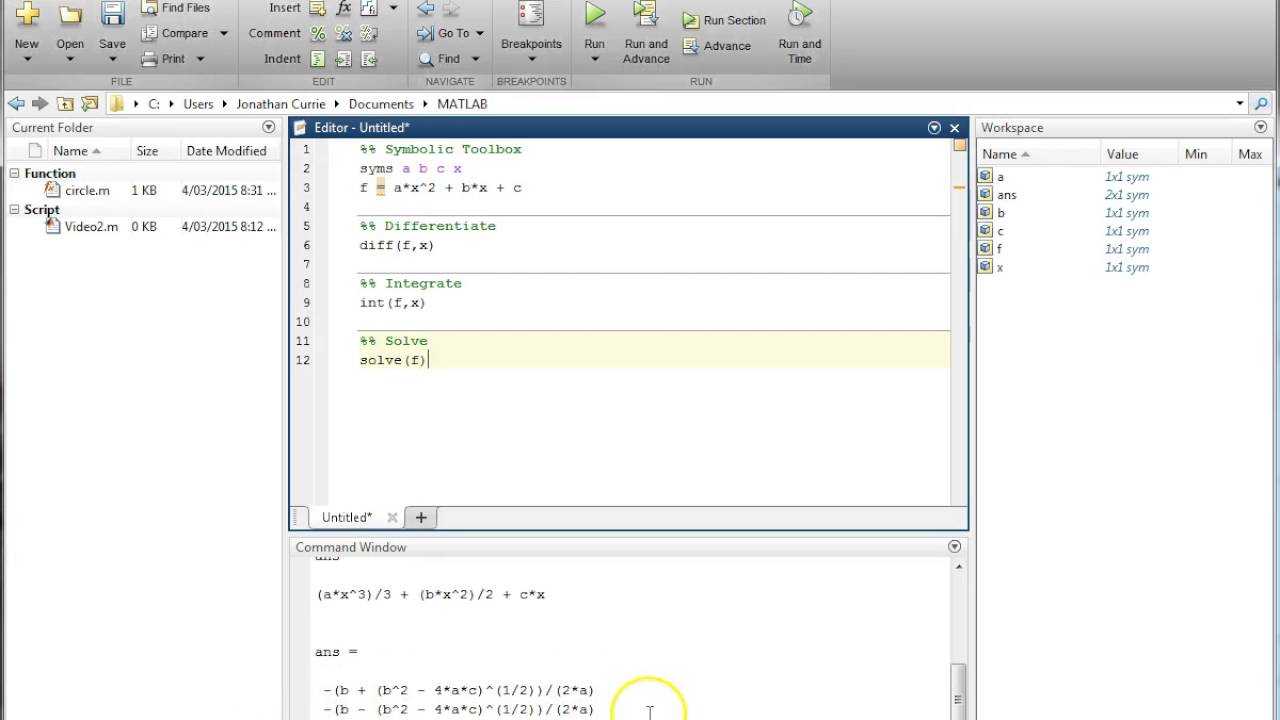
To visualize the representation, we can arrange the stimuli according to response-pattern dissimilarity, such that stimuli are placed close together if they elicited similar response patterns and far apart if they elicited dissimilar response patterns. Each cell of the matrix then compares the response patterns elicited by two images and the matrix is symmetric about a diagonal of zeros. These dissimilarities for all pairs of stimuli are assembled in the representational dissimilarity matrix (RDM). The dissimilarity between two patterns can be measured as 1 minus the correlation (0 for perfect correlation, 1 for no correlation, 2 for perfect anticorrelation). 
An RDM is a square symmetric matrix, each entry referring to the dissimilarity between the activity patterns associated with two stimuli (or experimental conditions).įigure 1 Computation of a representational dissimilarity matrix. For each pair of experimental stimuli, the response patterns elicited in a brain region or model representation are compared to determine the stimuli’s representational dissimilarity. RSA characterizes the representation in each brain region by a representational dissimilarity matrix (RDM Figure 1). It goes beyond testing for information in regional response patterns and enables us to handle condition-rich experiments without predefined stimulus categories, to test conceptual and computational models, and to relate representations between humans and monkeys (Kriegeskorte et al. RSA is a particular versatile version of MVPA. Representational Similarity Analysis (RSA) In the middle ground, a roaming “searchlight” can be used to examine the whole brain one voxel at a time, hence providing specificity in localisation while still using local pattern information. In others, the pattern of the whole brain is classified. In some cases, one can test regions of interestes if a priori reason can be established. Many different classifiers have been used to build the correspondence between activity pattern and mental state, including linear-discriminant analysis and support vector machines. Such information of orientation was lost in traditional fMRI analyses. For example, MVPA has been used to reveal sensitivity to orientation in V1 region in human brains. In achieving this, pattern classifiers are often used to relate distinct patterns of activation within the brain to corresponding mental states. However, increasing number of studies have shown that fine-grained information can also be obtained from the fMRI signal at a scale that is smaller than this. 
Traditional neuroimaging analysis techniques are designed to detect regional “activation” in the brain, which is often at centimetre scale. Cambridge Clinical Research Centre for Affective Disorders (C2AD).Cambridge Centre for Ageing & Neuroscience (CamCAN).Centre for Attention, Learning and Memory (CALM).V3.Using cognitive theory and innovations in neuroscience to understand and improve mental wellbeing across the lifespan Fleming, Creation andĪnalysis of biochemical constraint-based models: the COBRA Toolbox Stalidzans, Alejandro Maass, Santosh Vempala, Michael Hucka, MichaelĪ. Scott Hinton, William A.īryant, Francisco J. Le, Ding Ma, Yuekai Sun, Lin Wang, James T. Valcarcel, Inigo Apaolaza, Susan Ghaderi, MasoudĪhookhosh, Marouen Ben Guebila, Andrejs Kostromins, Nicolas Sauls,Īlberto Noronha, Aarash Bordbar, Benjamin Cousins, Diana C. Keating, Vanja Vlasov, Stefania Magnusdottir,Ĭhiam Yu Ng, German Preciat, Alise Zagare, Siu H.J. Mendoza, Anne Richelle, Almut Heinken, Hulda S. Laurent Heirendt, Sylvain Arreckx, Thomas Pfau, Sebastian N. It is widely used for modelling, analysing and predicting a variety of metabolic phenotypes using genome-scale biochemical networks. It implements a comprehensive collection of basic and advanced modelling methods, including reconstruction and model generation as well as biased and unbiased model-driven analysis methods. The COnstraint-Based Reconstruction and Analysis Toolbox is a MATLAB software suite for quantitative prediction of cellular and multicellular biochemical networks with constraint-based modelling. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
